An alternative standardization is scaling features to lie between a given minimum and maximum value, often between zero and one, or so that the maximum absolute value of each feature is scaled to unit size. This can be achieved using MinMaxScaler or MaxAbsScaler, respectively.
The motivation to use this scaling include robustness to very small standard deviations of features and preserving zero entries in sparse data. and the standard deviation is 1.
The values of each feature in a dataset can vary between random values. So, sometimes it is important to scale them so that this becomes a level playing field. Through this statistical procedure, it's possible to compare identical variables belonging to different distributions and different variables.
Let's see how to scale data in Python:
1. Let's start by defining the data_scaler variable:
data_scaler = preprocessing.MinMaxScaler(feature_range=(0, 1))
2. Now we will use the fit_transform() method, which fits the data and then transforms it (we will use the same data as in the previous recipe):
data_scaled = data_scaler.fit_transform(data)
A NumPy array of a specific shape is returned. To understand how this function has transformed data, we display the minimum and maximum of each column in the array.
3. First, for the starting data and then for the processed data:
print("Min: ",data.min(axis=0))
print("Max: ",data.max(axis=0))
The following results are returned:
Min: [ 0. -1.5 -1.9 -5.4] Max: [3. 4. 2. 2.1]
4. Now, let's do the same for the scaled data using the following code:
print("Min: ",data_scaled.min(axis=0))
print("Max: ",data_scaled.max(axis=0))
The following results are returned:
Min: [0. 0. 0. 0.] Max: [1. 1. 1. 1.]
After scaling, all the feature values range between the specified values.
To display the scaled array, we will use the following code:
[[1. 0. 1. 0. ] [0. 1. 0.41025641 1. ] [0.33333333 0.87272727 0. 0.14666667]]
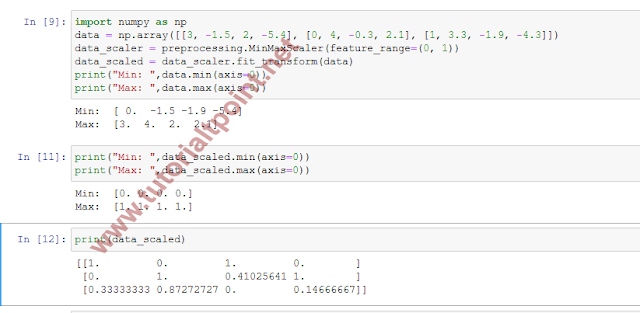
Here is the complete example

0 comments :
Post a Comment
Note: only a member of this blog may post a comment.